Articles
Article Tools
Stats or Metrics
Article
Original Article
Exp Neurobiol 2018; 27(5): 350-364
Published online October 31, 2018
https://doi.org/10.5607/en.2018.27.5.350
© The Korean Society for Brain and Neural Sciences
iPSC Modeling of Presenilin1 Mutation in Alzheimer's Disease with Cerebellar Ataxia
Ling Li1,†, Jee Hoon Roh2,†, Eun Hyuk Chang3,4, Yoonkyung Lee1, Suji Lee1, Minchul Kim1, Wonyoung Koh1, Jong Wook Chang3,4, Hee Jin Kim3,5, Mahito Nakanishi6, Roger A. Barker7, Duk L. Na3,5*, and Jihwan Song1*
1CHA Stem Cell Institute, Department of Biomedical Science, CHA University, Seongnam 13488, Korea.
2Department of Neurology, Asan Medical Center, University of Ulsan College of Medicine, Seoul 05505, Korea.
3Neuroscience Center, Samsung Medical Center, Seoul 06351, Korea.
4Stem Cell & Regenerative Medicine Institute, Samsung Medical Center, Seoul 06351, Korea.
5Department of Neurology, Samsung Medical Center, Seoul 06351, Korea.
6Research Center for Stem Cell Engineering, National Institute of Advanced Industrial Science and Technology (AIST), Ibaraki 305-8561, Japan.
7John van Geest Cambridge Centre for Brain Repair, Department of Clinical Neuroscience, University of Cambridge, Cambridge CB2 0PY, UK.
Correspondence to: *To whom correspondence should be addressed.
Jihwan Song, TEL: 82-31-881-7140, FAX: 82-31-881-7249
e-mail: jsong5873@gmail.com
Duk L. Na, TEL: 82-2-3410-3591, FAX: 82-2-3410-0052
e-mail: dukna@naver.com
†These authors contributed equally to the work.
Abstract
Disease modeling of Alzheimer's disease (AD) has been hampered by the lack of suitable cellular models while animal models are mainly based on the overexpression of AD-related genes which often results in an overemphasis of certain pathways and is also confounded by aging. In this study, we therefore developed and used induced pluripotent stem cell (iPSC) lines from a middle-aged AD patient with a known presenilin 1 (PSEN1) mutation (Glu120Lys; PS1-E120K) and as a control, an elderly normal subject. Using this approach, we demonstrated that the extracellular accumulation of Aβ was dramatically increased in PS1-E120K iPSC-derived neurons compared with the control iPSC line. PS1-E120K iPSC-derived neurons also exhibited high levels of phosphorylated tau, as well as mitochondrial abnormalities and defective autophagy. Given that the effect of aging is lost with iPSC generation, these abnormal cellular features are therefore indicative of PSEN1-associated AD pathogenesis rather than primary changes associated with aging. Taken together, this iPSC-based approach of AD modeling can now be used to better understand AD pathogenesis as well as a tool for drug discovery.
Graphical Abstract

Keywords: Alzheimer disease, stem cell, Presenilin1, Amyloid beta, Tau, Autophagy
INTRODUCTION
To date, the great majority of disease modifying clinical trials in Alzheimer's disease (AD) have not succeeded despite promising preclinical data. The potential reasons for this failure is that; (1) the agents so trialled are aimed at the wrong pathological target; (2) there is a loss of target engagement; (3) the treatment is given too late in the disease course; or (4) the previous models are affected by the effects of aging beyond AD pathogenesis [1,2].
Conventional AD models are mainly based on the overexpression of AD-related genes which overemphasizes by its very nature certain pathogenic pathways, thereby limiting the clinical translation of such approaches. In particular, the most popular mouse models of AD are based on the overexpression of each or combinations of amyloid precursor protein (
Using this approcah, we have demonstrated that the level of extracellular deposition of Aβ dramatically increased in PS1-E120K iPSC-derived neurons, compared with the control iPSC-derived neurons. Furthermore, we showed that PS1-E120K iPSC-derived neurons expressed high levels of phosphorylated tau along with abnormal mitochondrial activity and defective autophagy.
MATERIALS AND METHODS
The patient met the criteria for AD as recommended by the National Institute on Aging-Alzheimer's Association [6] and the normal elderly subject fulfilled the criteria of a normal elderly control as defined by Christensen [7]. We performed detailed neuropsychological tests, MRI, [18F]-Florbetaben or [11C]-Pittsburgh compound B (PIB) amyloid PET scans, and blood draw for iPSC generation from each participant. The Institutional review board of CHA University (1044308-201612-BR-031-02), Samsung Medical Center (2013-03-087), and Asan Medical Center (2015-0699) approved the study protocol, and informed written consent was obtained from each participant. All methods were performed in accordance with the relevant guidelines and regulations.
Mononuclear cells (MNCs) were isolated freshly from the peripheral blood of the AD patient and normal subject using the Ficoll-Paque™ PLUS method (GE Healthcare, USA). Isolated peripheral-derived MNCs (PBMCs) were cultured for 4 days in MNC media containing 50 ng/ml interleukin-6 (IL-6), 50 ng/ml stem cell factor (SCF), 10 ng/ml thrombopoietin (TPO), 20 ng/ml Flt3 ligand (Flt-3L), 20 ng/ml interleukin-3 (IL-3), and 10 ng/ml granulocyte colony-stimulating factor (G-CSF) (all from WAKO, Japan) in StemFit AK03 medium (kindly provided by Ajinomoto, Japan). The MNCs were then infected with SeVdp (KOSM) 302L at MOI of 3~10 [8] and transferred into 6-well dishes coated with iMatrix-511 (Matrixome, Japan). From the next day, 500 ul of StemFit AK03 medium was added every day for 4 days, after which the medium was fully changed every other day until iPSC-like colonies emerged. Sub-culturing and expansion of the cells was then undertaken until the generated iPSCs became stable for characterization and storage, normally at passage 10.
Karyotyping was performed using a GTG-banding analysis (Korea Research of Animal Chromosomes, Korea). The iPSC pellets were incubated in lysis buffer containing 100 mM Tris-HCl (pH 8.0), 50 mM EDTA, 0.2% SDS, 200 mM NACL and 200 ug/ml Proteinase K for 3 hr at 55℃. Genomic DNA was isolated by isopropyl alcohol precipitation, followed by a 70% ethanol washing step. Genotyping of the PS1-E120K single nucleotide mutation was performed by PCR amplification of isolated iPSC genomic DNA, followed by DNA sequencing (Cosmo Genetech, Korea). The PS1 gene was amplified by PCR using the following primers (forward primer: GTA GAA TCT ATA CCC CAT TC; reverse primer: TCA CCT TAT AGC ACC TGT AT).
Total RNAs were manually isolated using the TRIzol reagent (Life Technologies) lysis and isopropyl alcohol precipitation. Complementary DNAs (cDNAs) were synthesized using the cDNA synthesis Kit (Cosmo Genetech, Korea). RT-PCR amplification was done in a final volume of 20 ul containing 200 ng/ul cDNA for each sample. Primers were used as follows: GAPDH (forward primer: TGA CCA CAG TCC ATG CCA TCA CTG C; reverse primer: GTC ATA CCA GGA AAT GAG CTT GAC A); OCT4 (forward primer: CTG AAG CAG AAG AGG ATC AC; reverse primer: GAC CAC ATC CTT CTC GAG CC); NANOG (forward primer: TTC TTG ACT GGG ACC TTG TC; reverse primer: GCT TGC CTT GCT TTG AAG CA); SOX2 (forward primer: GCT GCA AAA GAG AAC ACC AA; reverse primer: CTT CCT GCA AAG CTC CTA CC); LIN28 (forward primer: CAC CAT GGG CTC CGT GTC CAA CCA GCA G; reverse primer: TCA ATT CTG TGC CTC CGG GAG CAG GGT AGG). Q-PCR analysis was performed using TB Green™ Premix Ex Taq™ (TaKaRa) with 100 ng/ul cDNA for each sample. Primers were used as follows: PSEN1 (forward primer: ACA GAG TTA CCT GCA CCG TTG T; reverse primer: GCT ATA AGG TCA TCC ATG CCT G); DRP1 (forward primer: ATG TTG CAT CTG GAG GTG GT; reverse primer: CGT GAA CCT GCT AGA TGT GC); Fis1 (forward primer: CAG TTT GAG TAC GCC TGG TG; reverse primer: CCG GCT CAA GGA ATA CGA GA); Mfn1 (forward primer: CTG GCT GTC TTG TAC GTG TG; reverse primer: CAA GGT GAA TGA GCG GCT TT); Mfn2 (forward primer: GAC CCC GTT ACC ACA GAA GA; reverse primer: GCT CTG GGA CAA AGT TCT GC).
After total RNA isolation and cDNA synthesis, Sendai virus detection RT-PCR was performed using a final volume of 20 ul with a 200 ng/ul concentration for each sample. Primers were used as follows: SeVdp NP (forward primer: AGA CCC TAA GAG GAC GAA GAC AGA; reverse primer: ACT CCC ATG GCG TAA CTC CAT AG) for detecting the nucleoprotein (NP) of Sendai virus vector (SeVdp) [9].
After genomic DNA isolation as described above, we detected the mycoplasma contamination in cell culture using the PCR-based e-Myco™ VALiD Mycoplasma detection Kit (iNtRON, Korea). PCR was done using a final volume of 20 ul containing 100 ng/ul of each sample [9,10].
Human iPSCs were harvested using a 0.5% TrypLE Select (ThermoFisher) treatment and used for embryoid body (EB) formation. Detached cells were transferred into 96-well Nunclon™ Sphera™ Microplate (ThermoFisher) that includes DMEM/F12 medium containing 20% Knockout serum replacement (KSR; Life Technology), 0.1 M nonessential amino acids (NEAA), 0.1 M 2-mercaptoethanol (Life Technologies) and 0.1 antibiotic-antimycotic solutions (ThermoFisher). The medium was changed every other day and the EBs were plated onto 0.2% gelatin-coated glass coverslips at day 8. EBs were then allowed to differentiate spontaneously for an additional 8 days.
For the cortical neuronal differentiation, iPSCs were dissociated into single cells and quickly re-aggregated in a 96-well Nunclon™ Sphera™ Microplate (ThermoFisher). The EBs were cultured in DMEM/F12 containing 20% knockout serum replacement (KSR), 0.1 M nonessential amino acids (NEAA), 0.1 M 2-mercaptoethanol, 0.1% antibiotic-antimycotic solutions (all from ThermoFisher), 10 uM SB431542 (Reagent Direct) and 10 uM LDN 193189 (Reagent Direct) for neural induction for 8 days. These neural induced EBs were dissociated using Accutase™ (Stem Cell Technologies) and were seeded onto 15 ug/ml poly-L-ornithine (Sigma) and 5 ug/ml laminin (Sigma)-coated dishes. Once neural precursor cells (NPCs) were formed, they were cultured in DMEM/F12 media containing 0.1 M nonessential amino acids (NEAA), 0.1 M 2-mercaptoethanol, 0.1 antibiotic-antimycotic solutions, 1% D-Glucose (Life Technologies), 1% B27 supplement without vitamin A (all from ThermoFisher), 200 mM L-Glutamine (Sigma), and 20 ng/ml basic fibroblast growth factor (bFGF; PeproTech Korea). Finally, NPCs were differentiated into cortical neurons for 10 weeks in Neurobasal A medium containing 1% B27 supplement without vitamin A, 1x Glutamax (all from ThermoFisher), 10 ng/ml brain-derived neurotrophic factor (BDNF), 10 ng/ml glial cell-derived neurotrophic factor (GDNF), 10 ng/ml nurotrophin-3 (NT3) (all from PeproTech Korea) on PLO/laminin-coated dishes [10].
The cells were fixed with 4% paraformaldehyde (PFA) for 15 min at room temperature (RT). Fixed cells were permeabilized and washed with TPBS containing 0.1% TritonX-100 (Sigma) in phosphate-buffered saline (PBS) and blocked with 5% normal horse serum (Vector Labs) for 30 min at RT. Afterwards, they were incubated with primary antibodies in blocking solution overnight with gentle rocking at 4℃. After washing three times with TPBS, they were incubated with secondary antibodies (ThermoFisher) for 90 min at RT, and then were counterstained using DAPI for 15 min. All fluorescence images were captured by confocal microscopy (TCSSP5II, Leica). The following primary antibodies were used: anti-OCT4 (1:200, Santa Cruz), anti-SOX2 (1:200, Millipore), anti-NANOG (1:200, R&D Systems), anti-SSEA-4 (1:100, Developmental Studies Hybridoma Bank)), anti-TRA-1-81 (1:100, CHEMICON), Tuj1 anti-tubulin beta III isoform (1:200, Millipore), anti-SMA (1:100, DAKO); anti-AFP (1:100, DAKO), anti-Nestin (1:200, R&D Systems), anti-Musashi (1:200, Millipore), anti-Map2 (1:200, Millipore), anti-TBR1 (1:100, Abcam), anti-CTIP2 (1:100, Abcam), 6E10 anti-Amyloid β (1:500, Covance), AT8 anti-p-tau (1:1000, ThermoFisher), anti-ChAT (1:200, Millipore) and anti-LC3B (1:500, Cell Signaling).
Conditioned media (CM) were collected from cultured neuronal cells (1×105) at 48 hours after the last medium change from 6 and 10 weeks of differentiation. The pellets of iPSC-derived neurons were resuspended in RIPA buffer (Biosesang) containing protease inhibitors (ThermoFisher) and phosphatase inhibitors (ThermoFisher). Then they were chilled on ice for 30 min and sonicated. The supernatants were collected after centrifugation at 13,000 rpm for 30 min to remove membrane lipids. Protein concentration was determined using a BCA Protein Assay Kit (ThermoFisher). Intracellular Aβ42 and Aβ40 were measured in a total of 1 µg proteins from 10 week-differentiated neurons. Aβ40 and Aβ42 levels were measured using the human Aβ40 and Aβ42 ELISA Kit according to the manufacture's instruction (IBL). ELISA plate reader (BioTek) was used to quantify Aβ40 and Aβ42 levels.
Cultured neurons in 6-well plates were harvested at 10 weeks of differentiation as cell pellets and were resuspended in RIPA buffer (BIOSESANG) containing protease inhibitors (ThermoFisher) and phosphatase inhibitors (ThermoFisher). They were then chilled on ice for 30 min and sonicated. The supernatants were collected after centrifugation at 13,000 rpm for 30 min to remove membrane lipids. Protein concentration was determined using a BCA Protein Assay Kit (ThermoFisher), and 25 ug of protein for each sample was electrophoresed on 8% or 12% gels. The separated samples were transferred onto a PVDF membrane and incubated with target antibodies. Protein bands were visualized by Immobilon Western (Millipore) and were detected using Chemi-Doc (Bio-Rad). The following primary antibodies were used: Tau5 anti-tau (1:1,000, ThermoFisher), AT8 anti-p-tau (1:1,000, ThermoFisher), anti-OPA1 (1:1,000, Santa Cruz), anti-Mfn1 (1:1,000, Abcam), anti-Mfn2 (1:1,000, Cell Signaling), anti-Drp1 (1:1,000, Cell Signaling), anti-Fis1 (1:1,000, Santa Cruz), anti-Ub (1:4,000, Santa Cruz), anti-LC3B (1:1,000, Cell Signaling), anti-LAMP2 (1:1,000, Santa Cruz), anti-Beclin1 (1:1,000, Cell Signaling), p62 anti-SQSTM1 (1:1,000, Santa Cruz), 4G8 anti-APP (1:500, Covance), anti-p-Drp1 (Ser637, 1:1000, Cell Signaling) and anti-β-actin (1:10000, Santa Cruz).
For autophagy flux assay, we treated cells with 10 uM chloroquine (CQ) for 12 h, a compound known to interference the lysosomal function [11]. After 12 h, we used the CytoID® Green Autophagy Detection Kit (Enzo Life Sciences Inc., Cat. No, ENZ-51031-K200) to detect the autophagic vesicles in live cells. Induction of autophagic flux can be visualized by enhanced accumulation of autophagic vesicles when lysosomal function is inhibited. CytoID green was co-stained with DAPI and the fluorescence images were captured using a confocal microscopy (TCSSP5II, Leica). The quantification for the number and size of autophgic vesicles were performed using Image J (particle analysis plugin).
All statistical analyses were performed using a one-factor analysis of variance (ANOVA) followed by a Fisher's LSD (Least Significant Difference) using the Statistical Analysis System (Enterprise 4.1, SAS Korea, Seoul, Korea). Significance was accepted at the 95% probability level. Data in graphs are presented as mean±SEM. p-value<0.05 (*), p-value<0.01 (**) and p-value<0.001 (***).
RESULTS
The patient was a 38-year-old woman who visited the outpatient clinic of Samsung Medical Center, Seoul, Korea. She had progressive memory impairment over a course of five years, which eventually resulted in disturbance of her daily life. She had 16 years of education and had no history of medical illness. A mildly spastic gait was noticed by her husband 6 months prior to presentation. Neurological examination revealed cerebellar dysfunction, including limb and truncal ataxia, dysdiadochokinesia, cerebellar dysarthria, and ocular abnormalities, such as overshooting saccades and saccadic intrusions during smooth pursuit eye movements. She also had bilateral spasticity in her legs and bradykinesia in both upper extremities. She demonstrated global cognitive problems with deficits in verbal and visual memory, language, and visuospatial functions with relative sparing of attention and frontal-executive function (Table 1).
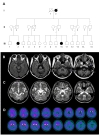
The patient had a family history of dementia and ataxia (Fig. 1A). Her grandmother (I:2 in Fig. 1A) had developed a late-onset dementia during the seventh decade of life and died ten years after the diagnosis. Her cousin (III:2 in Fig. 1A) was diagnosed with late-onset cerebellar ataxia at the age of 54 and died at 62 years of age due to aspiration pneumonia without evidence of cognitive impairment. Brain MR images of her cousin (III:2) demonstrated severe cerebellar atrophy (Fig. 1B) without prominent atrophy of the medial temporal lobes. Brain MRI of our patient revealed generalized cortical atrophy along with atrophy in the bilateral medial temporal lobes and cerebellum (Fig. 1C). Laboratory studies of the patient, including 24-hour urine copper levels were unremarkable. Given the tentative diagnosis of dementia associated with a familial cerebellar ataxia, genetic evaluations for spinocerebellar ataxia (SCA, 1, 2, 3, 6, 7, 8, 12, 17, 27), dentatorubral-pallidoluysian atrophy (DRPLA), and late-onset Friedreich's ataxia (Frataxin gene) were performed and shown to be negative. Huntingtin gene test was negative. Given the diagnosis of early onset dementia with spastic paraparesis, a genetic work-up for a Presenilin mutation was performed [12], and a mutation in the
Pittsburgh compound-B (PiB) positron emission tomography (PET) imaging of the patient revealed a deposition of Aβ in the striatum as well as in the bilateral frontal lobes, posterior cingulate cortices, and precuneus (Fig. 1D). These PiB-PET image findings of the patient are compatible with previous reports of familial AD with spastic paraparesis caused by
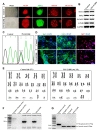
Isolated MNCs were reprogrammed using Sendai virus vector (SeVdp) which expresses four reprogramming factors (OCT3/4, SOX2, cMYC, and KLF4) [9]. In our iPSC generation procedure, we normally pick more than three individual clones and select the best growing clone among them for further analyses, including neuronal differentiation experiments. Although the effects of PS1-E120K mutation on Aβ deposition have been studied in the brains of familial AD patients previously [16,17], the pathological features of the patients are less known at cellular levels. For this reason, we generated an iPSC line from AD patient carrying a mutation for PS1-E120K using iPSC technology for the first time. Control and PS1-E120K mutation iPSC lines exhibited the typical expression of undifferentiated pluripotent stem cell markers, such as OCT4, SOX2, SSEA4 and TRA-1-81 (Fig. 2A, B). Genotyping of the established iPSC lines was confirmed using a conventional sequencing method (Fig. 2C). The differentiation potential of iPSC lines was assessed
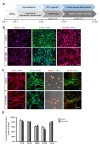
To characterize the cortical neurons derived from each iPSC line, we developed a modified cortical neuron differentiation protocol, based on previously published procedures (Fig. 3A) [10]. Differentiated cells expressed general neuronal markers (
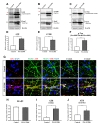
To analyze the functional aspects of familial AD (PS1-E120K), we investigated Aβ secretion in the conditioned medium (CM) from the iPSC-derived neurons at 6 weeks and 10 weeks after differentiation. We measured Aβ40 and Aβ42 levels 48 hr after the last medium change. Strikingly, PS1-E120K iPSC-derived neurons exhibited a dramatic increase in Aβ42 levels (over 2-fold) compared with the control iPSC-derived neurons at both 6 weeks and 10 weeks after differentiation (Fig. 4A). However, no difference in Aβ40 levels was detected (Fig. 4B). Importantly, the ratio of Aβ42/Aβ40 was significantly increased (over 2-fold) in PS1-E120K iPSC-derived neurons (Fig. 4C) compared with the control iPSC-derived line. Furthermore, intracellular Aβ42 levels and the ratio of Aβ42/Aβ40 was also significantly increased in PS1-E120K iPSC-derived neurons (Fig. 4D~F). We also observed a marked increase in APP expression levels in PS1-E120K iPSC-derived neurons (Fig. 4H and I). To detect the Aβ deposits, iPSC-derived neurons were stained with the anti-Aβ antibody 6E10 and 4G8 at 10 weeks of differentiation. Using a confocal microscopy we found a distinctive increase of extracellular Aβ deposits in PS1-E120K iPSC-derived neurons compared with the control iPSC-derived neurons (Fig. 4G, H). Given the difference in ages when the blood was drawn to generate both iPSC-derived neurons, the current findings suggest that the accumulation of Aβ protein can be driven by the PS1-E120K mutation beyond the effect of aging.
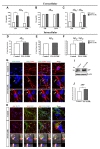
Tau protein normally exists as a soluble form in axons. However, as it becomes hyperphosphorylated, as in AD, it accumulates in dendrites and cell bodies [16,17]. To analyze the phosphorylated tau (p-tau) levels in the iPSC-derived neurons, we performed western blot analyses using antibodies against AT180 (phosphorylated at Thr231), p-Tau (phosphorylated at Ser400, Thr403/Ser404), and Tau5 (total tau). Then, we found that there exist multi-bands indicative of fractions in AT8, AT180 and p-Tau and the protein expression of each was strongly increased in the PS1-E120K iPSC-derived neurons compared with the control lines (Fig. 5A~F). These findings are in line with previous studies demonstrating that brain tissues from AD patients with severe Braak stages [18,19] and neural cells with PSEN1 and APP mutation showed multiple bands of p-tau (>50 kDa) compared with the controls [20]. We also found that there was an increase of total tau (Tau5) levels, which likely correlates with a robust increase in the aggregated tau fractions [20].
As we found the increase of fragmented tau in our PS1-E120K iPSC-derived neurons at 10 weeks of differentiation, we conducted tau ELISA to detect soluble tau using conditioned medium. We found that PS1-E120K patient exhibited significant increases in soluble tau both at 6 and 10 weeks of differentiation (data not shown). To further investigate this p-tau accumulation at the cellular level, we performed immunocytochemical analysis using an antibody against AT8 and found that, unlike in the control iPSC-derived neurons, AT8 expression in the PS1-E120K iPSC-derived neurons was localized to the cell bodies (Fig. 5G~J). Taken together, these results show that there are high levels of p-tau (AT8) which localizes to the cell bodies in the PS1-E120K iPSC-derived neurons, compared with the control iPSC-derived neurons.
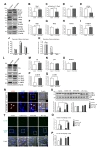
Mitochondrial dysfunction is a prominent feature in AD [22,23]. Recent studies have reported that extracellular Aβ and phosphorylated tau could be attributed to the impaired balance of mitochondrial fission and fusion [6,24]. To investigate whether mitochondria fission and fusion was impaired in our PS1-E120K iPSC-derived neurons, we measured the expression levels of mitochondria fission-related proteins, Drp1 (dynamin-related protein 1), p-Drp1 (for the detection of inhibitory phosphorylation of Drp1 at Ser637) and Fis1 (mitochondrial fission 1 protein); and fusion-related protein OPA1 (optic atrophy 1 gene protein), Mfn1 (membrane proteins mitofusin 1) and Mfn2 (membrane proteins mitofusin 2). Western blot analysis revealed a significant increase of Drp1 expression, a decrease of inhibitory p-Drp1 (Ser637) as well as the ratio of p-Drp1 (Ser637)/Drp1 and a slight increase of Fis1 in the PS1-E120K group compared with the control group (Fig. 6A~D). Furthermore, we found that the expression of OPA1, Mfn1 and Mfn2 was dramatically reduced in PS1-E120K group when compared with the control group (Fig. 6A, E, F and G). Q-PCR analyses revealed dramatic increases of fission-related gene (DRP1) expression and decrease of fusion-related gene (Mfn1) expression in the PS1-E12K iPSC-derived neurons compared with the control neurons (Fig. 6J and K). In addition, mitophagy-related proteins PINK1 and PARKIN showed dramatic increase in the PS1-E120K cells compared with the control lines (Fig. 6A, H and I). These results strongly suggest that
Defective autophagy function is known also to occur in AD which correlates with the presence of neuritic plaques and filamentous tau [25,26]. It was previously shown that PS1 is necessary for the degradation of abnormal proteins using autophagylysosome system [27]. To investigate this further in our neurons, we measured the expression levels of autophagy-related proteins, including Ubiquitin (Ub), p62 (SQSTM1, sequestosome 1; cargo protein marker), Beclin1 (autophagosome membrane formation-related protein), LC3b (light chain 3; autophagosome formation marker) and LAMP2 (lysosome-associated membrane proteins). Western blot analysis revealed that the expression levels of Ub and LC3b-II (active form of LC3b), but not p62, were significantly increased in PS1-E120K iPSC-derived neurons (Fig. 6L, M and P). To further investigate LC3B expression at the cellular level, we performed immunocytochemical analysis and found that PS1-E120K iPSC-derived neurons exhibit a dramatic increase of LC3b in the periphery of soma area, compared with the control group (Fig. 6R). In contrast, the chaperone-mediated autophagosome and lysosome membrane marker, LAMP2 showed a dramatic decrease in the PS1-E120K iPSC-derived neurons (Fig. 6L and Q). To further investigate the dysfunctioning autophagy, we performed autophagy flux assay with treatment of 10 uM chloroquine (CQ), a compound known to interfere with the lysosomal function. Autophagy flux assay showed a significant increase in LC3B-II expression after the CQ treatment in the PS1-E120K iPSC-derived NPC compared to the control (Fig. 6S). In addition, we performed the live cell staining with CytoID. In line with previous findings, the PS1-E120K iPSC-derived NPCs exhibited dramatic increases in the number of autophagic vesicles compared with the control cells (Fig. 6 T~P). Taken together, these results strongly suggest that
DISCUSSION
To better understand AD pathogenesis, we have utilized iPSC technology and established two iPSC lines, one harboring PS1-E120K mutation and one from a normal control. We did not use an isogenic control for two reasons; (i) the technical difficulty of doing this in the context of this specific mutation and (ii) the desire to look at aged control cells to exclude aging effects as the main pathological component of any disease pathogenic pathway that we identified in our mutation cell line.
We found that neurons so derived from the mutant cell line had the following characteristics. First, cortical neurons expressing TBR1+ and CTIP2+ and cholinergic neurons expressing ChAT+ were differentiated from both iPSC lines with no obvious differences in the differentiation propensity of the cell lines, nor any major morphological differences in the neurons so derived from. Second, the PS1-E120K iPSC-derived neurons exhibited higher levels of Aβ42, an altered Aβ42/Aβ40 ratio, and Aβ deposition compared with the iPSC-derived neurons developed from a normal elderly subject. Third, the levels of phosphorylated tau and its deposition in dendrites and cell bodies was seen only in the PS1-E120K iPSC-derived neurons. In line with previous studies demonstrated that increased Aβ42/40 ratio can lead increase in levels of p-Tau through GSK3β activation in AD patient iPSC-derived neurons [20,21], we deduced that PS1-E120K mutation can result in increased Aβ42/40 ratio and subsequent tau phosphorylation through the GSK3β activation. We have observed the same findings in another iPSC line generated from a patient carrying a different PSEN1 mutation (S170F), which exhibited a very similar pathophysiology as in the PS1-E120K iPSC line (data not shown). This new line also exhibited a very similar AD-related phenotypes, including increased levels of Aβ and p-Tau (AT8); impaired balance of mitochondrial fission and fusion; and abnormal expression of autophagy-related proteins. Given that similar AD pathophysiology can be observed from two independent PSEN1 iPSC lines, it will be likely that the results obtained from PS1-E120K iPSC line can truly represent the pathophysiology of AD patients carrying PSEN1 mutations.
There have been several recent studies on the establishment and characterization of PSEN mutant iPSC lines, but they are mainly focused on the increase of Aβ and Tau [26,28]. However, the present study investigated on not only Aβ and Tau levels but also mitochondria and autophagy defects. Mitochondria is a highly dynamic organelle and the intracellular population at any one time derives from a balance between fission and fusion [22]. Western blot analysis revealed that fission associated markers were increased, whereas fusion related markers were markedly decreased in PS1-E120K iPSC-derived neurons compared with the control iPSC-derived neurons. As extracellular Aβ and phosphorylated tau are implicated in mitochondrial fission and fusion in humans [8,24], we speculated that the increased secretion and aggregation of Aβ and phosphorylated forms of tau protein caused this imbalance of mitochondrial fission and fusion. Because mitochondrial fission is associated with autophagy and the genesis and degradation of autophagosomes [29], we thought the increase in mitochondrial fission may affect the clearance of pathologic protein accumulation through this pathway and this is what we found. Finally, given that the iPSC lines we generated were from a young donor, the changes in function of mitochondira and autophagy that we see must relate to the mutation and not an old cell.
There are many reports suggesting the possibility of defective autophagy in AD. It has been demonstrated that autophagy vacuoles (AVs) accumulate in human AD brains and are related to the presence of neuritic plaques and filamentous tau [25,28]. Furthermore, PS1 knockout mice showed a dysfunction of autophagy-lysosome system, including increase of LC3B and decrease of LAMP2 [27]. In this study, we observed significantly increased Ub and active forms of LC3b-II in the PS1-E120K iPSC-derived neurons, which strongly suggests that PS1-E120K mutation can induce impaired function of lysosome and the subsequent accumulation of AVs. In addition to the role of overproduction of Aβ through APP cleavage,
It should be stressed that there are several limitations to our study. First, the pateint and the elder sister of the patient had cerebellar atrophy and cerebellar symptoms and signs. Nevertheless, we were not able to further differentiate iPSC-derived neurons into Purkinje cells to evaluate relevant changes. Second, although the patient has a family history of dementia and cerebellar ataxia, the pedigree did not fit to an autosomal dominant patterns of inheritance. Thus, the patient may have a
In summary, we have shown that a
Figures
Tables
| Raw score | Percentile score | |
|---|---|---|
| Digit span Forward | 9/9 | 95.82 |
| Digit span Backward | 4/8 | 59.87 |
| K-BNT | 38/60 | 8.08 |
| RCFT Copy score | 29/36 | <0.01 |
| SVLT Immediate recall | 11/36 | 1.04 |
| SVLT Delayed recall | 0/12 | 0.04 |
| SVLT Recognition | 11/24 | <0.01 |
| RCFT Immediate recall | 2.5/36 | 3.67 |
| RCFT Delayed recall | 2.5/36 | 0.84 |
| RCFT Recognition | 13/24 | 0.13 |
| COWAT Animal | 10/20 | 0.62 |
| COWAT Supermarket | 10/20 | 14.23 |
| COWAT Phonemic | 28/45 | 87.29 |
| Stroop color reading | 81/112 | 28.1 |
K-BNT, Korean version of the Boston Naming Test; RCFT, Rey-Osterrieth Complex Figure Test; SVLT, Seoul Verbal Learning Test; COWAT, Controlled Oral Word Association Test.
References
- Dubois B, Hampel H, Feldman HH, Scheltens P, Aisen P, Andrieu S, Bakardjian H, Benali H, Bertram L, Blennow K, Broich K, Cavedo E, Crutch S, Dartigues JF, Duyckaerts C, Epelbaum S, Frisoni GB, Gauthier S, Genthon R, Gouw AA, Habert MO, Holtzman DM, Kivipelto M, Lista S, Molinuevo JL, O'bryant SE, Rabinovici GD, Rowe C, Salloway S, Schneider LS, Sperling R, Teichmann M, Carrillo MC, Cummings J, Jack CR. Preclinical Alzheimer's disease: definition, natural history, and diagnostic criteria. Alzheimers Dement 2016;12:292-323.
- Holtzman DM, Goate A, Kelly J, Sperling R. Mapping the road forward in Alzheimer's disease. Sci Transl Med 2011;3:114ps48.
- Götz J, Ittner LM. Animal models of Alzheimer's disease and frontotemporal dementia. Nat Rev Neurosci 2008;9:532-544.
- LaFerla FM, Green KN. Animal models of Alzheimer disease. Cold Spring Harb Perspect Med 2012;2:a006320.
- Mertens J, Paquola AC, Ku M, Hatch E, Böhnke L, Ladjevardi S, McGrath S, Campbell B, Lee H, Herdy JR, Gonçalves JT, Toda T, Kim Y, Winkler J, Yao J, Hetzer MW, Gage FH. Directly reprogrammed human neurons retain aging-associated transcriptomic signatures and reveal age-related nucleocytoplasmic defects. Cell Stem Cell 2015;17:705-718.
- McKhann GM, Knopman DS, Chertkow H, Hyman BT, Jack CR, Kawas CH, Klunk WE, Koroshetz WJ, Manly JJ, Mayeux R, Mohs RC, Morris JC, Rossor MN, Scheltens P, Carrillo MC, Thies B, Weintraub S, Phelps CH. The diagnosis of dementia due to Alzheimer's disease: recommendations from the National Institute on Aging-Alzheimer's Association workgroups on diagnostic guidelines for Alzheimer's disease. Alzheimers Dement 2011;7:263-269.
- Christensen KJ, Moye J, Armson RR, Kern TM. Health screening and random recruitment for cognitive aging research. Psychol Aging 1992;7:204-208.
- Manczak M, Calkins MJ, Reddy PH. Impaired mitochondrial dynamics and abnormal interaction of amyloid beta with mitochondrial protein Drp1 in neurons from patients with Alzheimer's disease: implications for neuronal damage. Hum Mol Genet 2011;20:2495-2509.
- Itoh M, Kawagoe S, Okano HJ, Nakagawa H. Integration-free T cell-derived human induced pluripotent stem cells (iPSCs) from a patient with lymphedema-distichiasis syndrome (LDS) carrying an insertion-deletion complex mutation in the FOXC2 gene. Stem Cell Res 2016;16:611-613.
- Kondo T, Asai M, Tsukita K, Kutoku Y, Ohsawa Y, Sunada Y, Imamura K, Egawa N, Yahata N, Okita K, Takahashi K, Asaka I, Aoi T, Watanabe A, Watanabe K, Kadoya C, Nakano R, Watanabe D, Maruyama K, Hori O, Hibino S, Choshi T, Nakahata T, Hioki H, Kaneko T, Naitoh M, Yoshikawa K, Yamawaki S, Suzuki S, Hata R, Ueno S, Seki T, Kobayashi K, Toda T, Murakami K, Irie K, Klein WL, Mori H, Asada T, Takahashi R, Iwata N, Yamanaka S, Inoue H. Modeling Alzheimer's disease with iPSCs reveals stress phenotypes associated with intracellular Aβ and differential drug responsiveness. Cell Stem Cell 2013;12:487-496.
- Rossi M, Munarriz ER, Bartesaghi S, Milanese M, Dinsdale D, Guerra-Martin MA, Bampton ET, Glynn P, Bonanno G, Knight RA, Nicotera P, Melino G. Desmethylclomipramine induces the accumulation of autophagy markers by blocking autophagic flux. J Cell Sci 2009;122:3330-3339.
- Brooks WS, Kwok JB, Kril JJ, Broe GA, Blumbergs PC, Tannenberg AE, Lamont PJ, Hedges P, Schofield PR. Alzheimer's disease with spastic paraparesis and ‘cotton wool’ plaques: two pedigrees with PS-1 exon 9 deletions. Brain 2003;126:783-791.
- Koivunen J, Verkkoniemi A, Aalto S, Paetau A, Ahonen JP, Viitanen M, Någren K, Rokka J, Haaparanta M, Kalimo H, Rinne JO. PET amyloid ligand [11C]PIB uptake shows predominantly striatal increase in variant Alzheimer's disease. Brain 2008;131:1845-1853.
- Verkkoniemi A, Kalimo H, Paetau A, Somer M, Iwatsubo T, Hardy J, Haltia M. Variant Alzheimer disease with spastic paraparesis: neuropathological phenotype. J Neuropathol Exp Neurol 2001;60:483-492.
- Klunk WE, Price JC, Mathis CA, Tsopelas ND, Lopresti BJ, Ziolko SK, Bi W, Hoge JA, Cohen AD, Ikonomovic MD, Saxton JA, Snitz BE, Pollen DA, Moonis M, Lippa CF, Swearer JM, Johnson KA, Rentz DM, Fischman AJ, Aizenstein HJ, DeKosky ST. Amyloid deposition begins in the striatum of presenilin-1 mutation carriers from two unrelated pedigrees. J Neurosci 2007;27:6174-6184.
- Spillantini MG, Goedert M. Tau pathology and neurodegeneration. Lancet Neurol 2013;12:609-622.
- Zempel H, Mandelkow E. Lost after translation: missorting of Tau protein and consequences for Alzheimer disease. Trends Neurosci 2014;37:721-732.
- Manczak M, Reddy PH. Abnormal interaction between the mitochondrial fission protein Drp1 and hyperphosphorylated tau in Alzheimer's disease neurons: implications for mitochondrial dysfunction and neuronal damage. Hum Mol Genet 2012;21:2538-2547.
- Trojanowski JQ, Lee VM. The role of tau in Alzheimer's disease. Med Clin North Am 2002;86:615-627.
- Choi SH, Kim YH, Hebisch M, Sliwinski C, Lee S, D'Avanzo C, Chen H, Hooli B, Asselin C, Muffat J, Klee JB, Zhang C, Wainger BJ, Peitz M, Kovacs DM, Woolf CJ, Wagner SL, Tanzi RE, Kim DY. A three-dimensional human neural cell culture model of Alzheimer's disease. Nature 2014;515:274-278.
- Ochalek A, Mihalik B, Avci HX, Chandrasekaran A, Téglási A, Bock I, Giudice ML, Táncos Z, Molnár K, László L, Nielsen JE, Holst B, Freude K, Hyttel P, Kobolák J, Dinnyés A. Neurons derived from sporadic Alzheimer's disease iPSCs reveal elevated TAU hyperphosphorylation, increased amyloid levels, and GSK3B activation. Alzheimers Res Ther 2017;9:90.
- Burté F, Carelli V, Chinnery PF, Yu-Wai-Man P. Disturbed mitochondrial dynamics and neurodegenerative disorders. Nat Rev Neurol 2015;11:11-24.
- Wang X, Su B, Lee HG, Li X, Perry G, Smith MA, Zhu X. Impaired balance of mitochondrial fission and fusion in Alzheimer's disease. J Neurosci 2009;29:9090-9103.
- Manczak M, Reddy PH. Abnormal interaction of VDAC1 with amyloid beta and phosphorylated tau causes mitochondrial dysfunction in Alzheimer's disease. Hum Mol Genet 2012;21:5131-5146.
- Chesser AS, Pritchard SM, Johnson GV. Tau clearance mechanisms and their possible role in the pathogenesis of Alzheimer disease. Front Neurol 2013;4:122.
- Nixon RA, Yang DS. Autophagy and neuronal cell death in neurological disorders. Cold Spring Harb Perspect Biol 2012;4:a008839.
- Neely KM, Green KN, LaFerla FM. Presenilin is necessary for efficient proteolysis through the autophagy-lysosome system in a γ-secretase-independent manner. J Neurosci 2011;31:2781-2791.
- Nixon RA, Yang DS. Autophagy failure in Alzheimer's disease--locating the primary defect. Neurobiol Dis 2011;43:38-45.
- Youle RJ, van der Bliek AM. Mitochondrial fission, fusion, and stress. Science 2012;337:1062-1065.



