Articles
Article Tools
Stats or Metrics
Article
Original Article
Exp Neurobiol 2018; 27(6): 508-525
Published online December 28, 2018
https://doi.org/10.5607/en.2018.27.6.508
© The Korean Society for Brain and Neural Sciences
Astrocyte Specificity and Coverage of hGFAP-CreERT2 [Tg(GFAP-Cre/ERT2)13Kdmc] Mouse Line in Various Brain Regions
Yongmin Mason Park1,2,3,†, Heejung Chun2,3,†, Jeong-Im Shin1,2,3, and C. Justin Lee1,2,3*
1Division of Bio-Medical Science & Technology, Department of Neuroscience, KIST School, Korea University of Science and Technology, Seoul 02792, Korea.
2Center for Glia-Neuron Interaction, Korea Institute of Science and Technology (KIST), Seoul 02792, Korea.
3Center for Cognition and Sociality, Institute for Basic Science, Daejeon 34126, Korea.
Correspondence to: *To whom correspondence should be addressed.
TEL: 82-42-878-9150, FAX: 82-42-878-9151
e-mail: cjl@ibs.re.kr
†
These authors contributed equally to this work.
Abstract
Astrocyte is the most abundant cell type in the central nervous system and its importance has been increasingly recognized in the brain pathophysiology. To study
Graphical Abstract
Keywords: Astrocytes, Glial fibrillary astrocytic protein, Cre recombinase, Tamoxfien
INTRODUCTION
Astrocytes are increasingly recognized as an important cell type in the brain pathophysiology. Physiologically, astrocytes not only function at developmental stages by guiding neuronal migration [1] but also contribute to neuronal functions including taking up neurotransmitters through their transporters [2], sensing neuronal activity through many neurotransmitter receptors [3] and releasing gliotransmitters such as glutamate, GABA, ATP and D-serine [4,5,6,7]. Additionally, they also buffer extracellular potassium for proper neuronal firing [8]. At pathological conditions, including neurodegenerative disease, astrocytic functions are known to be altered [9]. For example, it was reported that glutamate transporter 1 (GLT1) levels are reduced [10] and tonic γ-aminobutyric acid (GABA) release is increased in animal models of Alzheimer's disease [11], which cause excitotoxicity and neuronal inhibition, respectively.
During these processes, astrocytic gene expression is also altered and involved in these functional alternation. For studying
hGFAP-CreERT2 mouse lines, which are the most popular transgenic mouse line for genetically targeting astrocyte, were made by several groups [16,17,18,21]. Two mouse lines which were initially developed each from Vaccarino group [Tg(GFAP-cre/ERT2) 505Fmv/J] (MGI: 3774167), Kirchhoff group [Tg(GFAP-cre/ERT2) 1Fki] (MGI: 4418665), and Baker group were very well characterized with detailed quantitative analysis of astrocyte specificity. However, the other mouse line which is from Ken McCarthy group [Tg(GFAP-cre/ERT2) 13Kdmc], (MGI:3712447) has not been well characterized in various brain regions, despite its frequent use with over 75 citations of the original paper [23,24,25]. In this regard, it is necessary to characterize the cell types and its exact proportion in various brain regions for accurate interpretation of results.
Based on this need, we set out to characterize [Tg(GFAP-cre/ERT2) 13Kdmc], which we refer to as hGFAP-CreERT2, for assessing its efficiency of targeting astrocyte in various brain regions. To quantitatively analyze Cre expression pattern by crossing with Ai14 mouse line whose floxed tdTomato fluorescence gene can be sensitively and brightly activated by Cre. In this double transgenic mouse line, we analyzed the population of tdTomato positive cells to examine the ‘astrocyte specificity’, indicating the proportion of astrocyte in the total number of Cre expressing cells, and ‘astrocyte coverage’, indicating the proportion of Cre expressing cells in the total number of astrocytes, of the hGFAP-CreERT2 mouse line depending on brain regions.
MATERIALS AND METHODS
To generate reporter mice for visualizing Cre expression, we made double transgenic mouse line in C57BL/6J strain by crossing Ai14 (Rosa-CAG-LSL-tdTomato-WPRE::deltaNeo) mouse with hGFAP-CreERT2 mouse (hGFAP-CreERT2×Ai14). The mice have been bred in suitable space mice cage (Thoren, Hazleton, USA) with freely accessing to food and water and kept on a 12-hour-light-dark cycle. All experimental steps described in this paper were performed in accordance with the institutional guidelines for experimental animal care and use of the Korea Institute of Science and Technology (KIST; Seoul, Korea).
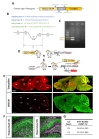
Mouse tails were digested overnight at 60℃, using 1 mg/ml Proteinase K (21560025-2, bioWORLD, Dublin, USA) in lysis reagent for genotyping (102-T, Viagenbiotech, Los Angeles, USA). On the following day, after inactivating the protein kinase K, the supernatant containing DNA was used for genotyping. The PCR reaction mixture contained Taq DNA Polymerase, MgCl2, dNTP mix (# QM13531, Bioquest, Seoul, Korea) and 1 µl genomic DNA template, 0.1 µM forward and reverse primer pairs and ddH2O. PCR was performed using the specific primer pairs as the detailed protocol is described in Fig. 1B. The reaction products were run on 1.5% agarose gels (HB0100500, E&S Bio Electronics, Daejeon, Korea) in TAE buffer (40 mM Tris: pH 7.6, 20 mM acetic acid and 1 mM EDTA) at 100 V and visualized using Nobel view (NOV001, Noble Bio, Hwaseong-si, Gyeonggi-do, Korea).
Stock solution of tamoxifen was made at concentration of 20 mg/ml in sunflower seed oil (Sigma, St.Louis, USA) mixture with 1/10 ethanol (64-17-5, Millipore, Burlington, USA). 100 µl of tamoxifen were administered into each hGFAP-CreERT2×Ai14 double transgenic mice (8~12weeks, male and female) by intraperitoneal (i.p.) injection per day for consecutive 5 days.
Mice were anesthetized with 2% avertin (20 µl/g, T48402, Sigma) by intraperitoneal injection and perfused with 4% paraformaldehyde accompanied by 0.1 M phosphate buffered saline (PBS) at room temperature. Extracted whole brains were post-fixed in 4% paraformaldehyde at 4℃ overnight. After that, post-fixed brains were incubated in 30% sucrose at 4℃ for more than 24 hours. Both sagittal and coronal sections were sliced with 30 µm thickness in cryostat microtome (Thermo Scientific, Waltham, USA). Sliced brain sections were washed three times with PBS and blocked with blocking solution (0.3% Triton X-100 (X-100, Sigma), 2% donkey serum (GTX27475, Genetex, Irvine, USA) and 2% goat serum (ab7481, Abcam, Cambridge, UK) in 0.1 M PBS for 1.5 hours. And then, these sections were incubated in suitable primary antibodies (rabbit anti-S100β antibody (ab41548, Abcam) and mouse anti-NeuN antibody (MAB377, Abcam) or chicken anti-tyrosine hydroxylase (P40101-0 TH, Abcam) or mouse anti-Calbindin antibody (C9848, Sigma) mixture with shaking at 4℃ overnight. After overnight incubation, brain sections were washed three times with PBS and incubated in matched secondary antibodies (Jackson, Bar Harbor, USA), they were also stained with 4′,6′-diamidino-2-phenylindole (DAPI, 62248, Pierce, Waltham, USA). Every sagittal sections' fluorescence images were taken by Operetta® High Contents Screening System. On the other hand, every coronal sections' fluorescence images were taken by Nikon A1 confocal microscope, and 30 µm Z stack images in 2-µm steps were processed by utilizing NIS-Elements (Nikon, Minato ku, Japan) software. Quantitative analysis method is described below paragraph. Reference atlases of mouse brain in the figure were obtained from Allen Brain Atlas (Reference Atlas, Version 2 (2011)) [26].
Coronal sections' fluorescence images were quantitatively analyzed by utilizing ImageJ (NIH) software. We identified astrocyte (also Bergmann glia) or neuron based on their morphology and co-localization with suitable each cell-type specific markers as we described above. After that, we counted and calculated Cre activated cell portion of each neurons and astrocytes which are co-localized with Cre mediated tdTomato fluorescence positive cell based on DAPI positive cell. Moreover, we defined ‘astrocyte specificity’ and ‘astrocyte coverage.’ The former indicates the percentage of S100β & tdTomato co-positive cells over total tdTomato positive cells, and the latter indicates that of S100β & tdTomato co-positive cells over total S100β positive cells.
RESULTS
According to the original description of hGFAP-CreERT2 mouse line, this mouse line was generated by micro injection of plasmid form of the transgene (2.2 kb human
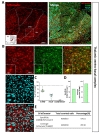
To characterize cell type specificity of tamoxifen-inducible Cre-expressing cells in hGFAP-CreERT2×Ai14 double transgenic mouse line, we administered tamoxifen or sunflower oil in these mice by i.p. injection for five consecutive days (Fig. 1D). Upon tamoxifen administration, we observed the tdTomato fluorescence in Cre-expressing cells. To scan and validate the Cre-expressing cells as an astrocyte, we performed immunohistochemistry by using an antibody against S100β as an astrocytic marker in sagittal brain sections (Fig. 1E, top). These sections showed ubiquitous Cre expression in all brain regions (Fig. 1E, top, left). Moreover, most of these cells showed co-localized immunoreactivity with S100β (Fig. 1E, top, right). In contrast, the control mouse (hGFAP-CreERT2×Ai14) with vehicle treatment (sunflower oil) showed minimal tdTomato fluorescence, indicating that there is virtually no leaky expression of Cre without tamoxifen (Fig. 1E, bottom, left). Furthermore, tdTomato fluorescence showed varying degree of co-localization with S100β in various brain regions, as evidenced by intensity of yellow color (Fig. 1E, top, right). For example, in cortex, hippocampus and cerebellum, tdTomato expressing cells were majorly merged with S100β showing intense yellow color, whereas in thalamus tdTomato expressing cells were rarely merged with S100β showing segregated two distinct colors (Fig. 1E, top, right). These results imply that the degree of astrocyte specificity is different in various brain regions.
These regional differences in the degree of astrocyte specificity could be originated from either the lack of specificity of S100β antibody to astrocytes or the region-dependent differential activity of human
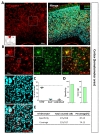
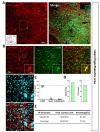
To investigate tamoxifen-inducible Cre-expressing cells in cortex, we performed immunohistochemistry by utilizing S100β as an astrocytic marker rather than GFAP, which is known to be differentially expressed in various brain regions [30]. We used NeuN (Neuronal nuclear antigen) as a neuronal marker for quantitative analysis (Fig. 2A). We found that tdTomato expressing cells in each image were predominantly co-localized with S100β in cortex (Fig. 2B and C, green, 87.37±1.45%). However, these tdTomato expressing cells were rarely co-localized with NeuN (Fig. 2B and C, cyan, 8.04±1.03%). Based on these high magnification images, we calculated ‘astrocyte specificity’ which is the ratio of the total counted number of tdTomato and S100β double positive (tdTomato+&S100β+) cells to that of tdTomato positive (tdTomato+) cells. Moreover, we also calculated ‘astrocyte coverage’ which is the ratio of the total counted number of tdTomato+&S100β+ cells to that of S100β+ cells. Based on these calculations, astrocyte specificity was 87.69% and coverage was 74.13% in cortex (Fig. 3D and E). These results indicate that hGFAP-CreERT2 mouse line has high astrocyte specificity with over 80% of specificity and high astrocyte coverage with over 70% of coverage in cortex.
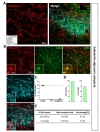
In striatum (caudoputamen) (Fig. 3A), tdTomato expressing cells in each image also showed pre-dominant S100β co-localization which was comparable or higher than that of cortex (Fig. 3B and C, green, 90.04±3.42%) and rare NeuN co-localization (Fig. 3B and C, cyan, 2.71±1.25%). Astrocyte specificity was 89.02% and coverage was 50.69%. (Fig. 3D and E). These results indicate that hGFAP-CreERT2 mouse line has high astrocyte specificity with over 80% of specificity, but low astrocyte coverage with under 70% of coverage in striatum.
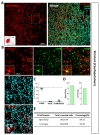
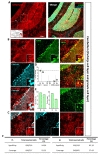
Next, we examined hippocampal CA1 layer (CA1) and Dentate Gyrus (DG). In hippocampal CA1 (Fig. 4A), tdTomato expressing cells in each image were highly co-localized with S100β (Fig. 4B and D, green, 92.61±1.49%) but not with NeuN (Fig. 4B and D, cyan, 5.21±2.10%). Astrocyte specificity was 92.83% and coverage was 56.28% (Fig. 4E and F, left).
In DG area (Fig. 4B), tdTomato expressing cells in each image were highly co-localized with S100β (Fig. 4C and D, green, 85.91±1.08%) than that of NeuN (Fig. 4C and D, cyan, 10.59±1.13%). Although, a few tdTomato expressing cells were found in subgranular zone where GFAP positive neuronal progenitor cells are known to be present [31], the total number was negligible (<5%). Astrocyte specificity was 86.13% and coverage was 61.44% (Fig. 4E and F, right). These results indicate that hGFAP-CreERT2 mouse line has high astrocyte specificity with over 80% of specificity, but relatively low astrocyte coverage with under 70% of coverage in both hippocampus sub-regions.
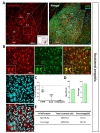
We further tested Cre expression pattern in lateral hypothalamus (LHA) (Fig. 5A). In LHA, tdTomato expressing cells were majorly S100β+ in each image (Fig. 5B and C, green, 92.05±2.39%), but NeuN+ cells were little (Fig. 5B and C, cyan, 5.48±2.61%). Astrocyte specificity was 92.20% and coverage was 79.24% in LHA (Fig. 5D and E). These results indicate hGFAP-CreERT2 mouse line has high astrocyte specificity with over 80% of specificity and high astrocyte coverage with over 70% of coverage in LHA.
Similar with LHA, we also tested in substantia nigra pars compacta (SNpC) by using S100β and tyrosine hydroxylase (TH) which marks dopaminergic neuron (Fig. 6A). Almost all of tdTomato expressing cells in SNpC were also S100β+ in each image (Fig. 6B and C, green, 97.74±0.59%), but no TH+ cells (Fig. 6B and C, cyan, 0.00%). In SNpC, astrocyte specificity was 97.64% and coverage was 72.94% (Fig. 6D and E). These results indicate that hGFAP-CreERT2 mouse line has the highest astrocyte specificity and high astrocyte coverage with over 70% of coverage in SNpC.
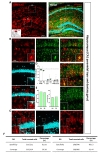
In addition, we also scrutinized Cre expression pattern in cerebellar Purkinje cell layer (PL) and granule cell layer (GL). Therefore, we stained with S100β for marking cerebellar astrocyte and Bergmann glia, and with Calbindin for marking Purkinje cell (Fig. 7A). In cerebellar PL, tdTomato expressing cells in each image were highly co-localized with S100β (Fig. 7B and D, green, 94.65±1.44%) but not Calbindin (Fig. 7B and D, cyan, 0.00%). Therefore, astrocyte specificity was 94.96% and coverage was 91.25% (Fig. 8E and F, left).
In GL area (Fig. 7B), tdTomato expressing cells in each image were much more co-localized with S100β+ cells (Fig. 7C and D, green, 83.46±1.91%) than that of Calbindin+ cells (Fig. 7C and D, cyan, 0.00%). Accordingly, astrocyte specificity was 83.29% and coverage was 57.69% in GL (Fig. 7E and F, right). These results indicate that hGFAP-CreERT2 mouse shows high astrocyte specificity with over 80% of specificity in both sub-regions of cerebellum. However, astrocyte coverage is different between these two sub-regions. Astrocyte coverage is high in PL with over 70% but low in GL with under 70%.
Finally, we characterized Cre expression in basolateral amygdala (BLA) and thalamic ventro-basal complex (VB) (Fig. 8A and 9A). Even though tdTomato expressing cells in each image were more co-localized with S100β, the ratio of co-localization with NeuN was also high (S100β+&tdTomato+: 66.28±2.80%, NeuN+&tdTomato+: 32.64±2.70%) (Fig. 8B and C, S100β: green, NeuN: cyan). As a result, astrocyte specificity was 65.34%, and coverage was 84.33% in BLA (Fig. 8D and E). These results indicate that hGFAP-CreERT2 mouse line has low astrocyte specificity with under 80% of specificity and high astrocyte coverage with over 70% of coverage in BLA.
Similarly, in thalamic VB, tdTomato expressing cells in each image showed non-specific expression between S100β+ and NeuN+ (S100β+&tdTomato+: 67.42±5.66%, NeuN+&tdTomato+: 32.33±5.48%) (Fig. 9B and C, S100β: green, NeuN: cyan). Based on this result, astrocyte specificity was 64.61%, and coverage was 89.30% in thalamic VB (Fig. 9D and E). These results indicate that hGFAP-CreERT2 mouse line has low astrocyte specificity with under 80% of specificity and high astrocyte coverage with over 70% of coverage in thalamic VB.
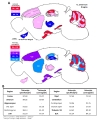
To sum up, we generated a color-coded brain map to represent the two quantitative aspects; ‘astrocyte specificity’ and ‘astrocyte coverage’ (Fig. 10A). Based on this brain map and table, astrocyte specificity of most of investigated regions was over 80% except in VB and BLA, whereas astrocyte coverage varied among the various brain regions ranging from 50% to 90% (Fig. 10A and B).
DISCUSSION
Our study quantitatively assessed the astrocyte specificity and coverage of hGFAP-CreERT2 mouse in various brain regions. We have demonstrated that this mouse line expressing Cre under the human
It is possible that this discrepancy could be due to the difference of mouse strain, too. The mouse lines from Kirchhoff group (FVB/N) and Baker group (FVB/NJ) are a different mouse strain from McCarthy group's mice line (C57BL/6J). A previous report has shown that FVB and C57BL/6N strains have different methylation activities [34]. Therefore, this different methylation condition could differentially regulate the human
Different tamoxifen administration protocol, including the types of injected ERT2 agonist or dosage of agonist, also can affect tamoxifen-mediated CreERT2 activation [20]. In previous reports, authors performed different tamoxifen injection protocol that is different from our study. For example, Vaccarino group used different ERT2 agonist, 4-hydroxytamoxifen (OHT, 33 mg/kg/day for one day) in mouse pups [17]. On the other hand, Baker group used the same ERT2 agonist, tamoxifen, but injected at different dosage of 225 mg/kg/day for five days [21]. However, Kirchhoff group used the same dosage of tamoxifen as in our study at 100 mg/kg/day for five days. Therefore, different tamoxifen administration methods can be a possible reason for the difference between the current study and previous reports.
The heterogeneity of astrocyte specificity in different brain regions, especially the significant neuronal expression in thalamic VB and BLA, might be caused by alternative gene silencing or heterogeneous activity of human
Our study provides very useful information regarding the optimal use of the hGFAP-CreERT2 mouse line, depending on the brain region of interest. For example, in cortex, LHA, SNpC and cerebellar Purkinje cell layer, the mouse line shows high astrocyte specificity with over 89% of specificity and high astrocyte coverage with over 70% of coverage. In those brain regions, this mouse line would be suitable for genetically modulating astrocytic gene expression in bidirectional ways, either gain-of-function or loss-of-function studies. In contrast, in striatum, hippocampus and cerebellar granule cell layer, the mouse line shows high astrocyte specificity with over 80% of specificity but covering only 50~60% of astrocytes. In those brain regions, this mouse line would be more suitable for genetically modulating gene expression in gain-of-function studies, rather than loss-of-function studies.
In conclusion, hGFAP-CreERT2 mouse line shows high astrocyte specific Cre expression in most of the brain regions. Our study demonstrates that this transgenic mouse line is very useful for manipulating genes of interest in astrocyte-specific manner, and perhaps is still one of the best genetic tools available for study of astrocyte function
Figures
References
- Kaneko N, Sawada M, Sawamoto K. Mechanisms of neuronal migration in the adult brain. J Neurochem 2017;141:835-847.
- Goubard V, Fino E, Venance L. Contribution of astrocytic glutamate and GABA uptake to corticostriatal information processing. J Physiol 2011;589:2301-2319.
- Porter JT, McCarthy KD. Astrocytic neurotransmitter receptors
in situ andin vivo . Prog Neurobiol 1997;51:439-455. - Beltrán-Castillo S, Olivares MJ, Contreras RA, Zúñiga G, Llona I, von Bernhardi R, Eugenín JL. D-serine released by astrocytes in brainstem regulates breathing response to CO2 levels. Nat Commun 2017;8:838.
- Guthrie PB, Knappenberger J, Segal M, Bennett MV, Charles AC, Kater SB. ATP released from astrocytes mediates glial calcium waves. J Neurosci 1999;19:520-528.
- Lee S, Yoon BE, Berglund K, Oh SJ, Park H, Shin HS, Augustine GJ, Lee CJ. Channel-mediated tonic GABA release from glia. Science 2010;330:790-796.
- Woo DH, Han KS, Shim JW, Yoon BE, Kim E, Bae JY, Oh SJ, Hwang EM, Marmorstein AD, Bae YC, Park JY, Lee CJ. TREK-1 and Best1 channels mediate fast and slow glutamate release in astrocytes upon GPCR activation. Cell 2012;151:25-40.
- Kofuji P, Newman EA. Potassium buffering in the central nervous system. Neuroscience 2004;129:1045-1056.
- Chun H, Lee CJ. Reactive astrocytes in Alzheimer's disease: a double-edged sword. Neurosci Res 2018;126:44-52.
- Guo Y, Duan W, Li Z, Huang J, Yin Y, Zhang K, Wang Q, Zhang Z, Li C. Decreased GLT-1 and increased SOD1 and HO-1 expression in astrocytes contribute to lumbar spinal cord vulnerability of SOD1-G93A transgenic mice. FEBS Lett 2010;584:1615-1622.
- Jo S, Yarishkin O, Hwang YJ, Chun YE, Park M, Woo DH, Bae JY, Kim T, Lee J, Chun H, Park HJ, Lee DY, Hong J, Kim HY, Oh SJ, Park SJ, Lee H, Yoon BE, Kim Y, Jeong Y, Shim I, Bae YC, Cho J, Kowall NW, Ryu H, Hwang E, Kim D, Lee CJ. GABA from reactive astrocytes impairs memory in mouse models of Alzheimer's disease. Nat Med 2014;20:886-896.
- Foo LC, Dougherty JD. Aldh1L1 is expressed by postnatal neural stem cells
in vivo . Glia 2013;61:1533-1541. - Zhuo L, Theis M, Alvarez-Maya I, Brenner M, Willecke K, Messing A. hGFAP-cre transgenic mice for manipulation of glial and neuronal function
in vivo . Genesis 2001;31:85-94. - Koh W, Park YM, Lee SE, Lee CJ. AAV-mediated astrocyte-specific gene expression under human
ALDH1L1 promoter in mouse thalamus. Exp Neurobiol 2017;26:350-361. - Casper KB, McCarthy KD. GFAP-positive progenitor cells produce neurons and oligodendrocytes throughout the CNS. Mol Cell Neurosci 2006;31:676-684.
- Casper KB, Jones K, McCarthy KD. Characterization of astrocyte-specific conditional knockouts. Genesis 2007;45:292-299.
- Ganat YM, Silbereis J, Cave C, Ngu H, Anderson GM, Ohkubo Y, Ment LR, Vaccarino FM. Early postnatal astroglial cells produce multilineage precursors and neural stem cells
in vivo . J Neurosci 2006;26:8609-8621. - Hirrlinger PG, Scheller A, Braun C, Hirrlinger J, Kirchhoff F. Temporal control of gene recombination in astrocytes by transgenic expression of the tamoxifen-inducible DNA recombinase variant CreERT2. Glia 2006;54:11-20.
- Slezak M, Göritz C, Niemiec A, Frisén J, Chambon P, Metzger D, Pfrieger FW. Transgenic mice for conditional gene manipulation in astroglial cells. Glia 2007;55:1565-1576.
- Srinivasan R, Lu TY, Chai H, Xu J, Huang BS, Golshani P, Coppola G, Khakh BS. New transgenic mouse lines for selectively targeting astrocytes and studying calcium signals in astrocyte processes
in situ andin vivo . Neuron 2016;92:1181-1195. - Chow LM, Zhang J, Baker SJ. Inducible Cre recombinase activity in mouse mature astrocytes and adult neural precursor cells. Transgenic Res 2008;17:919-928.
- Petryszak R, Keays M, Tang YA, Fonseca NA, Barrera E, Burdett T, Füllgrabe A, Fuentes AM, Jupp S, Koskinen S, Mannion O, Huerta L, Megy K, Snow C, Williams E, Barzine M, Hastings E, Weisser H, Wright J, Jaiswal P, Huber W, Choudhary J, Parkinson HE, Brazma A. Expression Atlas update--an integrated database of gene and protein expression in humans, animals and plants. Nucleic Acids Res 2016;44:D746-D752.
- Park H, Han KS, Seo J, Lee J, Dravid SM, Woo J, Chun H, Cho S, Bae JY, An H, Koh W, Yoon BE, Berlinguer-Palmini R, Mannaioni G, Traynelis SF, Bae YC, Choi SY, Lee CJ. Channel-mediated astrocytic glutamate modulates hippocampal synaptic plasticity by activating postsynaptic NMDA receptors. Mol Brain 2015;8:7.
- Petr GT, Sun Y, Frederick NM, Zhou Y, Dhamne SC, Hameed MQ, Miranda C, Bedoya EA, Fischer KD, Armsen W, Wang J, Danbolt NC, Rotenberg A, Aoki CJ, Rosenberg PA. Conditional deletion of the glutamate transporter GLT-1 reveals that astrocytic GLT-1 protects against fatal epilepsy while neuronal GLT-1 contributes significantly to glutamate uptake into synaptosomes. J Neurosci 2015;35:5187-5201.
- Sun XD, Li L, Liu F, Huang ZH, Bean JC, Jiao HF, Barik A, Kim SM, Wu H, Shen C, Tian Y, Lin TW, Bates R, Sathyamurthy A, Chen YJ, Yin DM, Xiong L, Lin HP, Hu JX, Li BM, Gao TM, Xiong WC, Mei L. Lrp4 in astrocytes modulates glutamatergic transmission. Nat Neurosci 2016;19:1010-1018.
- Dong HW. The allen reference atlas. A digital color brain atlas of the C57BL/6J male mouse. Chichester: Wiley, 2008.
- Deloulme JC, Raponi E, Gentil BJ, Bertacchi N, Marks A, Labourdette G, Baudier J. Nuclear expression of S100B in oligodendrocyte progenitor cells correlates with differentiation toward the oligodendroglial lineage and modulates oligodendrocytes maturation. Mol Cell Neurosci 2004;27:453-465.
- Hachem S, Aguirre A, Vives V, Marks A, Gallo V, Legraverend C. Spatial and temporal expression of S100B in cells of oligodendrocyte lineage. Glia 2005;51:81-97.
- Steiner J, Bernstein HG, Bielau H, Berndt A, Brisch R, Mawrin C, Keilhoff G, Bogerts B. Evidence for a wide extraastrocytic distribution of S100B in human brain. BMC Neurosci 2007;8:2.
- Kálmán M, Hajós F. Distribution of glial fibrillary acidic protein (GFAP)-immunoreactive astrocytes in the rat brain. I. Forebrain. Exp Brain Res 1989;78:147-163.
- Doetsch F, Caillé I, Lim DA, García-Verdugo JM, Alvarez-Buylla A. Subventricular zone astrocytes are neural stem cells in the adult mammalian brain. Cell 1999;97:703-716.
- Madisen L, Zwingman TA, Sunkin SM, Oh SW, Zariwala HA, Gu H, Ng LL, Palmiter RD, Hawrylycz MJ, Jones AR, Lein ES, Zeng H. A robust and high-throughput Cre reporting and characterization system for the whole mouse brain. Nat Neurosci 2010;13:133-140.
- Seymour PA, Sander M. Immunohistochemical detection of beta-galactosidase or green fluorescent protein on tissue sections. Methods Mol Biol 2007;411:13-23.
- Bai X, Saab AS, Huang W, Hoberg IK, Kirchhoff F, Scheller A. Genetic background affects human glial fibrillary acidic protein promoter activity. PLoS One 2013;8:e66873.
- Restrepo A, Smith CA, Agnihotri S, Shekarforoush M, Kongkham PN, Seol HJ, Northcott P, Rutka JT. Epigenetic regulation of glial fibrillary acidic protein by DNA methylation in human malignant gliomas. Neuro-oncol 2011;13:42-50.
- Garrick D, Fiering S, Martin DI, Whitelaw E. Repeat-induced gene silencing in mammals. Nat Genet 1998;18:56-59.
- Park D, Park SH, Ban YW, Kim YS, Park KC, Kim NS, Kim JK, Choi IY. A bioinformatics approach for identifying transgene insertion sites using whole genome sequencing data. BMC Biotechnol 2017;17:67.
- Morel L, Chiang MS, Higashimori H, Shoneye T, Iyer LK, Yelick J, Tai A, Yang Y. Molecular and functional properties of regional astrocytes in the adult brain. J Neurosci 2017;37:8706-8717.



